Translating information from DNA to proteins is the most expensive process a cell undertakes, and is central to all life. The basic principles of translation are simple - free floating ribosomes bind to mRNAs and translate the transcript one codon at a time, each of which is recognized by a corresponding tRNA. Although our understanding of how certain proteins and RNAs participate in translation has improved in recent years, we still lack a coherent view of what factors set the global pace of protein synthesis in a cell. To systematically integrate the various components of translation that have each been subjected to empirical measurements. Therefore to systematically study protein synthesis, we need high-throughput genomic tools that capture the translation dynamics at the level of an entire cell.
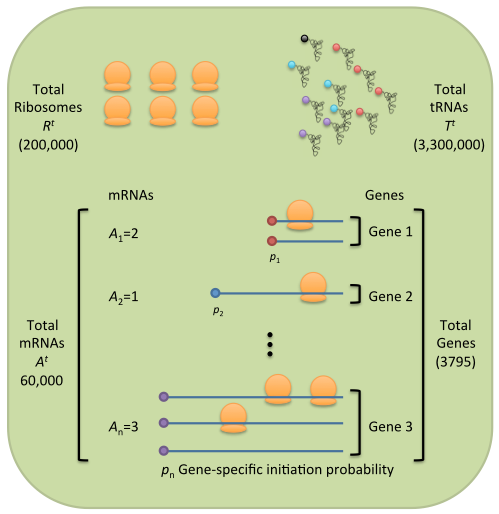
We have developed a mathematical and computational modeling framework that tracks every single ribosome, tRNA, and mRNA molecule within a cell defined by its biophysical parameters, and allows us to determine how translation of each gene is affected by all other genes. We also generate high-throughput ribosome-profiling datasets that provide detailed snapshots of translation by mapping each elongating ribosome in the cell. We employ this combined modeling/experimental approach to study how a cell modulates translation in different environments, including during diseases. By quantifying the dynamics of translation we study the mechanistic basis of adaptation in evolving populations and study the evolution of translational differences between homologous genes in divergent species.