Abstract
Patterns of diversification rate variation detected in phylogenetic hypotheses are frequently used to infer historical, ecological, and evolutionary processes. The parametric rate comparison (PRC) is a method for detecting rate variation in trees that models branch lengths as random variables drawn from familiar statistical distributions. iteRates is a library of functions for the R statistical computing environment for implementing PRC on phylogenetic trees. Here, we describe some of the functions in iteRates for subtree identification, tree manipulation, applying the PRC and K-clades PRC analyses, and conducting a whole-tree randomization test.
Software
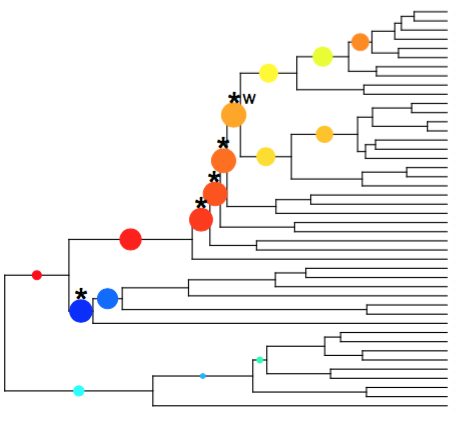
The library for parametric rate comparisons is released as an R package: iteRates with a detailed manual. To identify shifts in rates of diversification along lines of common ancestry, we iteratively compare each monophyletic subtree to the remainder tree obtained by pruning out the subtree.